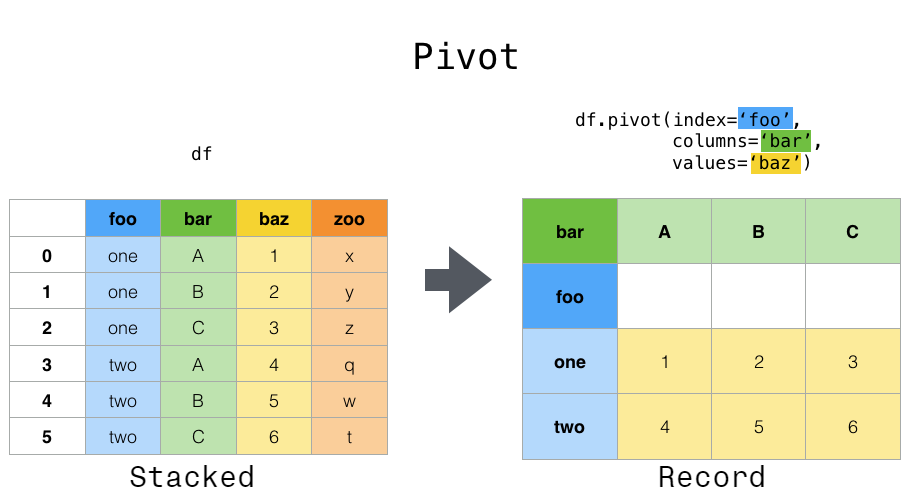
We’d need to create a new table where each row (the unit) is comprised of values of variables associated with each plot. What if instead of comparing records, we wanted to compare the different mean weight of each genus between plots? (Ignoring plot_type for simplicity). In surveys, the rows of surveys contain the values of variables associated with each record (the unit), values such as the weight or sex of each animal associated with each record. Here we examine the fourth rule: Each type of observational unit forms a table. Each type of observational unit forms a table.
#Stat transfer variables in rows not columns how to#
In the spreadsheet lesson, we discussed how to structure our data leading to the four rules defining a tidy dataset: To learn more about dplyr and tidyr after the workshop, you may want to check out this handy data transformation with dplyr cheatsheet and this one about tidyr.Īs before, we’ll read in our data using the read_csv() function from the tidyverse package readr. Moving back and forth between these formats is non-trivial, and tidyr gives you tools for this and more sophisticated data manipulation. Other times we want a data frame where each measurement type has its own column, and rows are instead more aggregated groups (e.g., a time period, an experimental unit like a plot or a batch number). For example, sometimes we want data sets where we have one row per measurement. The package tidyr addresses the common problem of wanting to reshape your data for plotting and usage by different R functions. The database connections essentially remove that limitation in that you can connect to a database of many hundreds of GB, conduct queries on it directly, and pull back into R only what you need for analysis. This addresses a common problem with R in that all operations are conducted in-memory and thus the amount of data you can work with is limited by available memory. The benefits of doing this are that the data can be managed natively in a relational database, queries can be conducted on that database, and only the results of the query are returned.

An additional feature is the ability to work directly with data stored in an external database. It is built to work directly with data frames, with many common tasks optimized by being written in a compiled language (C++). The package dplyr provides helper tools for the most common data manipulation tasks. Then, type library(tidyverse) to load the package. If you haven’t already done so, you can type install.packages("tidyverse") straight into the console. You should already have installed and loaded the tidyverse package. The existence of hidden arguments having default operations that new learners are not aware of.R expressions are used in a non standard way, which can be confusing for new learners.The results from a base R function sometimes depend on the type of data.The tidyverse package tries to address 3 common issues that arise when doing data analysis in R: The tidyverse package is an “umbrella-package” that installs tidyr, dplyr, and several other useful packages for data analysis, such as ggplot2, tibble, etc. It pairs nicely with tidyr which enables you to swiftly convert between different data formats for plotting and analysis.
dplyr is a package for helping with tabular data manipulation. Reshape a data frame from long to wide format and back with the spread and gather commands from the tidyr package.īracket subsetting is handy, but it can be cumbersome and difficult to read, especially for complicated operations.Describe the concept of a wide and a long table format and for which purpose those formats are useful.Use summarize, group_by, and count to split a data frame into groups of observations, apply summary statistics for each group, and then combine the results.Use the split-apply-combine concept for data analysis.Add new columns to a data frame that are functions of existing columns with mutate.Link the output of one dplyr function to the input of another function with the ‘pipe’ operator %>%.